Design and establishment of a Φ29-based replication system in vivo
Bacteriophage Φ29 is a lytic, double-stranded DNA bacteriophage belonging to the Salasmaviridae family33. First discovered in 1965 (ref. 34), it remains the smallest known phage that infects Bacillus. The Φ29 genome (19,282 bp) encodes 27 protein-coding genes and a prohead RNA that is essential for genome packaging (Fig. 1a)33. The encoded proteins are involved in replicating and packaging the genome and lysing the host cell33,35,36. The Φ29 genome is linear and contains origin of replication sequences (oriL and oriR) at each end. The genome is capped at its 5′ ends by covalently bound terminal proteins (TPs); these proteins prime replication by the Φ29 DNAP37,38. The Φ29 genome is an important model system for studying protein-primed DNA replication.
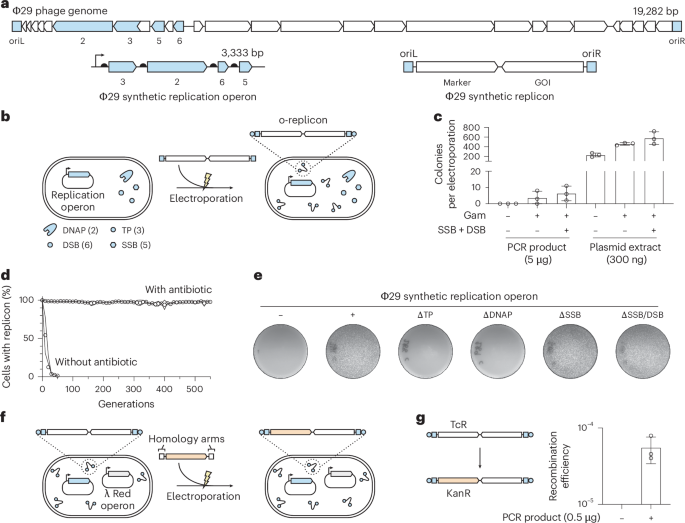
Fig. 1: Establishing and characterizing a Φ29-based replication system in E. coli. a, A synthetic replication operon was designed on the basis of genes 3, 2, 6 and 5, encoding the TP, DNAP, DSB and SSB, respectively. The synthetic replicon was designed to encode an antibiotic resistance gene and a gene of interest (GOI) and was flanked by the left and right origins of replication (oriL and oriR) derived from the Φ29 genome. b, Φ29 synthetic replicons could be established by electroporating a PCR-derived product into cells harboring the Φ29 synthetic replication operon on a single-copy plasmid. c, Efficiency of establishing Φ29 synthetic replicons by electroporation. A helper plasmid was used to additionally express gam and the genes encoding the Φ29 SSB and DSB. The Φ29 synthetic replicons extracted from cells could be transformed with a higher efficiency (n = 3 technical replicates—three electroporations into the same batch of electrocompetent cells); error bars indicate the s.d. We note that ‘Plasmid extract’ includes the replicon and copurified plasmid encoding the replication operon; thus, the quantity of replicon transformed is lower than 300 ng. d, Stability of the TcR–GFP Φ29 synthetic replicon over 550 generations with or without tetracycline, as assessed by maintenance of GFP fluorescence using flow cytometry (n = 4 biological replicates); lines indicate the s.d. e, Essentiality of genes in the synthetic replication operon for transformation of extracted Φ29 synthetic replicons. f, Established Φ29 synthetic replicons can be engineered using lambda Red recombination by the electroporation of a PCR amplicon flanked by homologies to the replicon. g, Efficiency of replacing the tetracycline resistance gene on the Φ29 synthetic replicon with a kanamycin resistance gene (n = 3 technical replicates—three electroporations into the same batch of electrocompetent cells); error bars indicate the s.d. Source data Full size image
We aimed to generate a synthetic system for the controlled replication of a linear Φ29 replicon in E. coli. Toward this goal, we designed a synthetic replication operon consisting of the genes encoding TP (gene 3), DNAP (gene 2), double-stranded DNA-binding protein (DSB; gene 6) and single-stranded DNA-binding protein (SSB; gene 5) (Fig. 1a); a combination of these proteins has been used to amplify portions of the Φ29 genome or heterologous genes in vitro39,40. We arranged these genes into an operon under the control of an IPTG-inducible promoter, PtacIPTG, on a single-copy plasmid (Fig. 1a). All genes were computationally codon-optimized for use in E. coli and their expression was driven by synthetic ribosome-binding site sequences41,42,43.
We designed a Φ29 synthetic replicon to be replicated by the synthetic replication operon; the replicon consisted of genes encoding tetracycline resistance (TcR) and GFP, flanked by the left and right Φ29 origins of replication (oriL and oriR) (Fig. 1a). We attempted to establish the Φ29 synthetic replicon in E. coli by electroporating 5 µg of the Φ29 synthetic replicon, generated by PCR amplification, into E. coli cells harboring the synthetic replication operon plasmid (Fig. 1b). However, this yielded no tetracycline-resistant colonies (Fig. 1c). This indicated that our initial attempt to establish the replicon system was unsuccessful.
We hypothesized that the establishment of the replicon may be limited by degradation of the electroporated PCR product in E. coli. To maximize the chance of establishing the synthetic replicon, we generated helper plasmids that express genes encoding the Gam protein (derived from the lambda phage) to inhibit host nucleases (RecBCD and SbcCD), as well as the Φ29 DSB and SSB to further protect the electroporated replicon DNA.
Upon electroporation of the Φ29 synthetic replicon into cells bearing the helper plasmids and the synthetic replication operon, we observed a small number of tetracycline-resistant colonies that exhibited green fluorescence. These experiments suggested that the Φ29 synthetic replicon was established in these cells (Fig. 1c); this conclusion was supported by additional genotyping and sequencing experiments (Supplementary Fig. 1) and by experiments where we could reduce the replicon copy number by downregulating expression from the replication operon by targeting the promoter driving the operon with dCas9 (Supplementary Fig. 2). Extracting the established replicons from cells bearing the helper plasmids and the synthetic replication operon and electroporating them into fresh cells enabled substantially improved transformation efficiencies, even when the helper plasmids providing genes encoding Gam and Φ29 DSB and SSB were not present in the recipient cells (Fig. 1c).
We measured the growth of cells with or without the synthetic Φ29 replication operon and the Φ29 synthetic replicon (Supplementary Fig. 3). This revealed the replication operon itself to lead to a mild reduction in cell fitness (doubling times of 49–52 min), whereas cells harboring both the replication operon and the Φ29 synthetic replicon exhibited substantially slower growth (doubling time of 87 min) and a lower maximum cell density (Supplementary Fig. 3).
The Φ29 synthetic replicon is stably inherited
To assess the stability of the Φ29 synthetic replicon through many rounds of cell division, we measured the proportion of cells that maintained the replicon, as indicated by GFP fluorescence, over 550 generations (Fig. 1d and Supplementary Fig. 4). Without tetracycline, the replicon was lost in less than 50 generations. In the presence of tetracycline, the replicon was stably maintained for the entire 550 generations tested. Therefore, the Φ29 synthetic replicon can be stably maintained for many generations, as required for continuous directed evolution experiments.
Defining the genes required for maintenance of the Φ29 synthetic replicon
Next, we defined the genes necessary for replication of the Φ29 synthetic replicon in E. coli. We generated a set of single-copy plasmids harboring variants of the Φ29 synthetic replication operon; in each member of the set, an individual gene (or pair of genes) was disrupted. We then electroporated the Φ29 synthetic replicon into cells containing each of these plasmids and assessed the formation of colonies that exhibited tetracycline resistance and green fluorescence. This experiment revealed that only two genes, encoding TP and DNAP, are essential for replicative maintenance of the Φ29 synthetic replicon (Fig. 1e). We passaged cells harboring the Φ29 synthetic replication operon, with or without the genes encoding the SSB and DSB proteins, and a TcR–GFP Φ29 synthetic replicon for 50 generations and assessed whether the Φ29 synthetic replicons remained intact by colony PCR (Supplementary Fig. 5). We only observed full-length replicons, suggesting that the absence of the SSB and DSB did not notably impair replication fidelity. Our results are in agreement with in vitro efforts to replicate the Φ29 genome using only TP and DNAP and the self-replication of TP-encoding and DNAP-encoding replicons in synthetic protocells44,45,46.
Efficient engineering of the Φ29 synthetic replicon
As the establishment of new Φ29 synthetic replicons from PCR amplicons was a low-efficiency process, we sought to develop a strategy for engineering replicons that were established in cells to introduce new DNA sequences of interest. The lambda Red recombination system, from the lambda phage, has been extensively used for engineering bacterial genomes and plasmids through the introduction of linear DNA flanked by short (>30 bp) stretches of homology to the target site47,48. To assess whether this recombination system could be used to engineer Φ29 synthetic replicons, we tested replacing the tetracycline resistance gene on the replicon with a kanamycin resistance gene. We electroporated 0.5 μg of a PCR product consisting of a kanamycin resistance gene flanked by 60-bp homologies into cells harboring a Φ29 synthetic replication operon plasmid, a tetracycline-resistance-conferring Φ29 synthetic replicon and a plasmid encoding the lambda Red recombination system components under arabinose-inducible control (Fig. 1f). These experiments gave rise to kanamycin-resistant colonies, where the tetracycline resistance gene was replaced, at a frequency approaching 10−4 (Fig. 1g). All modifications of the Φ29 synthetic replicon performed in this work were conducted using this approach.
Optimization of the Φ29 synthetic replication operon
Next, we sought to improve the performance of the designed Φ29 synthetic replication operon by experimentally optimizing DNA sequences that may control the relative levels of the proteins produced from the operon. To achieve this, we generated a Φ29 synthetic replication operon library in which we varied three regions of the designed operon: we randomized the spacer sequence between the Shine–Delgarno sequence and the start codon of each gene (this sequence can modulate the efficiency of translation initiation, which is rate determining for translation in E. coli), we randomized the first codon following the start codon (to allow for stability tuning according to the N-end rule, which differs between the Bacillus species that Φ29 phage infect and E. coli) and we randomized codons 2–7 to synonyms (as the sequence in this region has been shown to influence translational efficiency) (Fig. 2a)49,50,51,52,53.
Fig. 2: Improving Φ29 synthetic replication operon performance. a, A Φ29 synthetic replication operon library was generated where, for each gene in the operon, the spacer sequence between the Shine–Delgarno sequence and the start codon was randomized, the N-terminal codon following the start codon was saturated and the following six codons were synonymized. b, The Φ29-opt operon, integrated on a single-copy plasmid, supports a higher replicon copy number, as measured by GFP fluorescence and qPCR (n = 6 biological replicates for fluorescence; n = 3 biological replicates for qPCR). Fluorescence values were normalized to culture density. c, Extracted Φ29 synthetic replicons could be efficiently transformed into cells where the Φ29-opt operon was genomically integrated (n = 3 technical replicates—three electroporations into the same batch of electrocompetent cells). Error bars indicate the s.d. CFU, colony-forming unit. d, The Φ29-opt operon, encoded from a single-copy plasmid, enables improved growth when maintaining a TcR–GFP Φ29 synthetic replicon (n = 16 biological replicates). The boundaries indicate the s.e.m. Source data Full size image
We passaged cells transformed with the Φ29 synthetic replication operon library and a Φ29 synthetic replicon to enrich operons with favorable properties. To select for operons that supported increased replicon copy numbers, we picked colonies exhibiting stronger GFP fluorescence. We identified an operon, Φ29-opt (Supplementary Fig. 6), with improved performance; this operon supported a higher replicon copy number (Fig. 2b), improved transformation of extracted replicons (>106 transformants per µg of transformed replicon; Fig. 2c), improved establishment of new replicons (Supplementary Fig. 7) and faster growth while maintaining a Φ29 synthetic replicon (doubling time reduced from 87 to 66 min) to higher cell densities (Fig. 2d and Supplementary Fig. 4). We hypothesize that the improved growth is because of the increased capacity to maintain the Φ29 synthetic replicon, ensuring stable transfer of the replicon to daughter cells at every division.
We next tested whether Φ29-opt could support longer Φ29 synthetic replicons. We used lambda Red recombination to insert variable lengths of the CFTR gene and a CmR gene onto a KanR–GFP replicons to yield replicons with lengths of 6.6 and 10.5 kb. The longer replicons were established and maintained, as assessed by genotyping and sequencing (Supplementary Fig. 8).
Highly mutagenic Φ29-based orthogonal replication system
We generated Φ29 synthetic replication operons, based on Φ29-opt, with variants (N62D and F65S, alone or in combination) of the Φ29 DNAP; in in vitro experiments, these individual DNAP mutants are reported to mutate the DNA that they replicate while retaining strand-displacement activity54,55. We used these operon variants to demonstrate the creation of a highly mutagenic orthogonal replication system in E. coli (Fig. 3).
Fig. 3: The Φ29-based replication system is an orthogonal replication system in E. coli. a, An orthogonal replication system enables hypermutation of a target DNA sequence without interfering with the high-fidelity replication of genomic DNA. b, Error-prone mutants of the Φ29 DNAP do not increase the genomic mutation rate but do increase the replicon mutation rate. The mutagenic T7 DNAP mutant (mut_G) has previously been reported18. We used fluctuation analysis to calculate the mutation rate from the frequency of a TAG stop codon reversion in a genomically or replicon-integrated chloramphenicol resistance gene (Q38TAG) (n = 12 biological replicates). Error bars indicate the median ± upper or lower 95% bounds. Source data Full size image
To measure the extent to which these Φ29 DNAP mutants mutagenize the Φ29 synthetic replicon, we generated a replicon harboring a chloramphenicol resistance gene with an in-frame stop codon (Q38TAG). After ten generations of growth, with each DNAP variant and a wild-type (WT) control, we measured the fraction of chloramphenicol-resistant cells arising from a point mutation in the TAG stop codon to generate sense codons. These experiments allowed us to determine the apparent mutation rates of the replicon in the presence of each DNAP using fluctuation analysis56,57. This analysis revealed that the DNAP mutants were highly mutagenic on the replicon (Fig. 3 and Supplementary Fig. 9). The N62D;F65S double mutant outperformed the individual mutants, with a mutation rate approaching 10−4 substitutions per base, per generation on the replicon (Fig. 3). To determine the mutational spectra of these error-prone DNAP mutants, we grew cells harboring a Φ29 synthetic replicon for up to 50 generations and measured the accumulation of mutations using next-generation sequencing (Supplementary Fig. 10). We primarily observed C/G>A/T mutations, although all types of mutations were detected (Supplementary Figs. 10–12). We found mutation rates calculated from this sequencing data to be between 9.6 × 10−5 and 1.8 × 10−4 for the N62D;F65S DNAP double mutant (Supplementary Figs. 11 and 12). Moreover, the introduced mutations did not appear to exhibit positional bias (Supplementary Figs. 11 and 12).
Next, we integrated a chloramphenicol resistance gene containing an in-frame stop codon (Q38TAG) into the genome of E. coli. We introduced the Φ29 DNAP mutants into these cells and measured the fraction of chloramphenicol-resistant cells, arising from point mutations in the genomic TAG stop codon that generate a sense codon, after ten generations. These experiments allowed us to determine the apparent mutation rates of the genome in the presence of each DNAP using fluctuation analysis56,57. This analysis revealed that expression of the WT or mutant Φ29 DNAPs did not lead to changes in the apparent genomic mutation rate with respect to control cells without a Φ29 DNAP (Fig. 3). By contrast, expression of an error-prone mutant of T7 DNAP led to increased mutagenesis of the genome18,58. We additionally measured genomic mutation rates in the presence of a Φ29 replicon but did not observe a difference in the apparent mutation rates (Supplementary Fig. 13).
Overall, our data demonstrate that mutagenic Φ29 DNAPs enable mutation rates approaching 10−4 substitutions per base, per generation on the replicon, but do not measurably affect the mutation rate of the genome. We conclude that the Φ29-based replication system constitutes an orthogonal replication system in E. coli.
Continuous evolution of tigecycline, ceftazidime and cefotaxime resistance
Next, we sought to use the Φ29 orthogonal replication system to continuously evolve new phenotypes. We initially investigated evolving the tetracycline-resistance-conferring tetA gene into a gene that confers resistance to tigecycline. This evolution has been carried out using previously reported continuous evolution approaches9,16. We passaged cells containing a tetA–GFP replicon and the N62D;F65S error-prone DNAP in increasingly higher concentrations of tigecycline, until a final concentration of 40 µg ml−1 was reached (12 passages; Supplementary Fig. 14). The resultant pools of cells grew on plates containing tigecycline concentrations as high as 50 µg ml−1, whereas the pools of cells before continuous evolution grew on 0.2 µg ml−1 but not on 0.5 µg ml−1 (Fig. 4 and Supplementary Fig. 15). We sequenced replicons from cells growing on high tigecycline concentrations and found that certain mutations had convergently emerged in our three independent replicates (Supplementary Data 1). Of note are W233, which was substituted to C or S in every clone, A393, which was substituted to S or D in every clone, P193, which was substituted to S, T or N in 79% of clones, G388, which was substituted to V in some clones from all replicates but in all clones from the third replicate, and V355, which was substituted to F in two of the three replicates (Fig. 4a). We cloned select sequences from each replicate into a standard circular plasmid (colE1) to further validate the performance of these mutants (Supplementary Fig. 16). This revealed that all mutants tested could confer resistance to tigecycline concentrations between 10 and 30 µg ml−1, whereas the WT tetA control was unable to grow on 1 µg ml−1 tigecycline. We also tested the susceptibility of these evolved variants to counter-selection as mediated by a combination of fusaric acid and ZnCl 2 , revealing a range of sensitivities (Supplementary Fig. 17). The most sensitive variant had 103–104-fold higher sensitivity compared to the parent TetA and, thus, constitutes an improved dual positive–negative selection marker59.
Fig. 4: Continuous evolution of tigecycline and ceftazidime resistance using the Φ29-based replication system. a, Structure of TetA (AlphaFold prediction) with the residues that were substituted in at least two independent evolution replicates shown in stick representation. All observed substitutions are summarized in Supplementary Data 1. b, Pools of cells before and after continuous evolution for enhanced tigecycline resistance. Other pools are shown in Supplementary Fig. 15. Hits from the evolution were also cloned into a standard circular plasmid and validated (Supplementary Fig. 16). c, Structure of the TEM-1 β-lactamase (PDB 1XPB) with the residues that were substituted in at least two independent ceftazidime resistance evolution replicates shown in stick representation. All observed substitutions are summarized in Supplementary Data 2. d, Pools of cells before and after continuous evolution for enhanced ceftazidime resistance. Other pools are shown in Supplementary Fig. 19. Hits from the evolution were also cloned into a standard circular plasmid and validated (Supplementary Fig. 20). Full size image
Next, we sought to rapidly evolve TEM-1 β-lactamase for improved activity against the third-generation cephalosporin ceftazidime. Third-generation cephalosporin antibiotics differ from earlier generations because of their improved resistance to β-lactamase activity60. We passaged cells harboring a replicon encoding the TEM-1 β-lactamase in increasing concentrations of ceftazidime, ranging from 2 to 500 µg ml−1 (Supplementary Fig. 18). We completed three passages in 3 days, yielding pools of cells that grew on 500 µg ml−1 ceftazidime (Fig. 4 and Supplementary Fig. 19). The input pools of cells grew poorly on just 0.5 µg ml−1 ceftazidime and did not grow on 1 µg ml−1. We sequenced replicons from cells growing on high ceftazidime concentrations and found that certain mutations had convergently emerged in our four independent replicates (Supplementary Data 2). Of note are E102, which was substituted to K in every sequenced clone, R162, which was substituted to S or N in every sequenced clone, E237, which was substituted to K in some clones in two independent evolution replicates, and E166, which was substituted to D in some clones from three of the four independent evolution replicates (Fig. 4c). We cloned select sequences from each replicate into a standard circular plasmid (colE1) to further validate the performance of these mutants (Supplementary Fig. 20). This revealed that all mutants tested could confer resistance to ceftazidime concentrations between 200 and 500 µg ml−1, whereas the WT control was unable to grow on 1 µg ml−1 ceftazidime.
Subsequently, we assessed the possibility of evolving multi-resistant TEM-1 β-lactamase variants. We passaged the pools obtained from the ceftazidime resistance evolution in increasing concentrations of cefotaxime, ranging from 0.5 to 100 µg ml−1, while simultaneously selecting for both ceftazidime and carbenicillin resistance by supplementing these antibiotics at 100 µg ml−1 (Supplementary Fig. 21). We completed four passages in 4 days, yielding pools of cells that grew on 200 to 500 µg ml−1 cefotaxime while maintaining resistance to both carbenicillin and ceftazidime (Supplementary Fig. 21). By contrast, the ceftazidime-resistant pools, which were the input to this evolution, grew poorly or not at all at greater than 10 µg ml−1 of cefotaxime. We sequenced replicons from cells growing on high cefotaxime concentrations and found that certain mutations had convergently emerged between replicates 1 and 3 and between replicates 2 and 4 (Supplementary Fig. 21 and Supplementary Data 2). Replicates 2 and 4 had diverged most from the ceftazidime-resistant clones, with P165T, A170D and M180T substitutions in every replicon sequenced from these replicates, in addition to other substitutions. We cloned select sequences from each replicate into a standard circular plasmid (colE1) to further validate the performance of these mutants (Supplementary Fig. 21). This revealed that all mutants tested could confer resistance to cefotaxime concentrations between 50 and 200 µg ml−1, whereas clone 1 from the ceftazidime resistance evolution grew poorly on 10 µg ml−1 cefotaxime. All tested clones maintained ceftazidime and carbenicillin resistance (Supplementary Fig. 21). Given that these evolutions are run with multiple replicons per cell, it would in principle be possible for multiresistance to either emerge in a single genotype or for distinct resistances to be conferred by separate genotypes. In this case, all obtained clones fall into the former category. We hypothesize that the improved heritability associated with a single genotype conferring multiresistance may provide a selective advantage.
These experiments demonstrate that the Φ29 orthogonal replication system enables the rapid and reproducible evolution of new gene function.
— Source: Nature Biotechnology (https://www.nature.com/articles/s41587-025-02944-x)